An Entirely Synthetic Yeast Genome Is Nearly Complete
Posted on Categories Discover Magazine
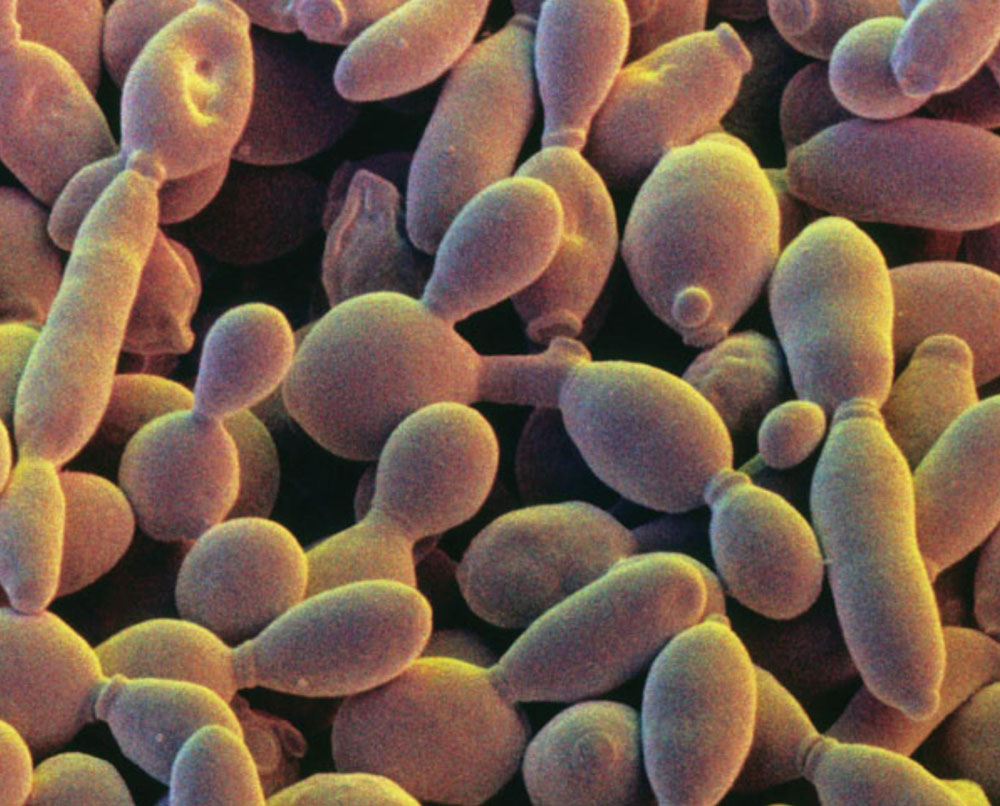
Scientists are five steps closer to synthesizing the entire genome of baker’s yeast, a feat that, once accomplished, will push the field of synthetic biology into a new frontier.
An international team of researchers led by NYU Langone geneticist Jef Boeke on Thursday announced it constructed and integrated five “designer” chromosomes into Saccharomyces cerevisiae. This collaboration, known as the Synthetic Yeast 2.0 project (Sc2.0), unveiled the first-ever “designer chromosome” back in 2014, which brings the official total of made-from-scratch chromosomes to 6 of baker’s yeast 16.
In other words, over 30 percent of a living organism’s genetic material can be substituted with artificial code, and it won’t be long before Sc2.0 researchers reach 100 percent—an entirely synthetic organism. Discover spoke with Boeke about the project’s recent success:
“We’re aiming to finish all 16 chromosomes by the end of this calendar year,” says Boeke. “There are several more that are already complete, and they are entering the debugging and characterization phase, which can take a good year.”
Once, or when, they get there, apart from building the first synthetic, complex organism, a new generation of geneticists will be able to entertain many of the most vexing questions about the structure and function of genomes. Customized yeast cells will churn out novel medicines, biofuels and other useful compounds with more precision and speed than ever before.
“We’re all really excited about seeing the end in sight, but the end is really just the beginning. Once we combine everything into one cell, that’s when the real fun begins,” Boeke says. “That’s when the power of a fully synthetic genome will become apparent.”
The breakthrough was outlined in a series of seven papers, with over 200 authors listed, published Thursday in the journal Science.
How’d They Do It?
Synthesizing a yeast chromosome is sort of like knitting a molecular sweater. It all starts with a plan: Via computer, scientists write DNA sequences, cutting “junk DNA” that doesn’t code for proteins, and “jumping DNA” that can drift into different locations and trigger unwanted mutations. They keep what works, and cut what doesn’t.
Next, biotech companies that specialize in DNA synthesis assemble short sequences based on the plan, which are then stitched together into longer strands. The strands are further weaved into even longer chunks. Finally, scientists integrate those chunks of DNA into yeast cells, replacing the natural chromosome with synthetic pieces until it reflects the design they made on the computer.
“The constraints changed from year to year due to improvements in DNA synthesis and assembly methods, and we had to keep up with the changes constantly,” Joel Bader, a project collaborator and professor of biomedical engineering at Johns Hopkins University, wrote in an email to Discover.
The synthetic chromosome isn’t an exact copy of its natural counterpart; it’s a bit leaner. In fact, Sc2.0 scientist have finished the planning stage for the remaining chromosomes, and the resulting Saccharomyces cerevisiae 2.0 genome plan has roughly 1 million nucleotide-level differences from the natural version.
Next Steps
Once all 16 chromosomes are successfully constructed, the next step will be slipping them into a single cell to see if they all work together. Leslie Mitchell, a post-doctoral fellow at Boeke’s NYU Lagone lab, has already designed a yeast strain that contains three synthetic chromosomes.
Considering there are so many unknowns in genetics, the chromosome construction system works surprisingly well.
“One of the mottos of synthetic biology is we hope to fail, because that’s when you learn,” says Boeke. “So, in a way, our failure rate is almost disappointingly low.”
Still, there were wrinkles that forced the team devise new approaches to iron them out and operate more efficiently. In doing so, they’ve established a technical foundation for GP-write, a related initiative that aims to synthesize sets of plant and human genomes within the decade.
Driving down the costs associated with synthesizing DNA and stitching sequences together remains a formidable obstacle if scientists hope to build more complex genomes. It cost $1.25 million for the entire yeast genome, not including materials, labor, sequence verification and debugging.
“The human genome is about 300 times the size of the yeast genome, and to mow through that efficiently is going to be a very costly exercise if we do it at 10 cents a base pair,” Boeke says. “We need to drive that price down to the point where the DNA is basically free.”
And although the human genome is a target for the GP-write project, we can withhold allusions to The Island of Doctor Moreau and the rise of strange, human-like hybrids. GP-write will hold its next big meeting May 9-10 in New York City, and Boeke said they’ll address the raging debate surrounding human germline editing.
“I draw the line at humans,” says Boeke. “For the GP-write project, we’re just not going to go there. We’re going to ask participants to just not go in that direction.”
What They’ll Do With It
When the yeast genome is complete, synthetic biology will enter a new age of inquiry and applications. What rules do genomes abide by? How much can you scramble a genome before it’s essentially a new species? How do genes interact with each other? How important is their position within the chromosome? These questions are all fair game, and scientists might get answers.
Boeke’s team designed a built-in scrambling system in the yeast genome, which isn’t possible with native yeast. It’s essentially high-speed evolution in a test tube. Scientists can generate millions of genome variations in a milliliter of water and identify the versions that provide the best performance for an intended outcome.
“Rather than doing experiments brute force, we have millions of cells in a test tube each doing a different experiment for us,” says Bader. “Then we ask the winners what they did by sequencing their DNA.”
For industrial purposes, this is incredibly useful. Genetically modified yeast already produce painkillers, spider silk, insulin, biofuels and, of course, alcoholic beverages. But the scrambling technique hones in on the most useful, efficient variants, which will locate the perfect yeast strain for a given task faster.
“You’re shuffling the genome like a deck of cards, but you can actually cheat,” says Boeke. “You can remove cards from the deck, or you can put ten copies of the ace of spades in the deck if you’d like.”
For Boeke, collaboration’s work represents the next logical step in the gradual evolution of genome engineering.
“We started out doing genome engineering using natural selection, and crossbreeding,” he says. “And then we entered the age of genetic engineering, doing things one gene at a time, and now we’re at the full-on genome stage of that evolution.”
Like Boeke said, the fun is just getting started.